For decades, the therapeutic antibody landscape has been heavily dominated by the Immunoglobulin G (IgG) isotype. Its long serum half-life, well-understood pharmacokinetics, and manageable structural properties have made it the gold standard for biopharmaceuticals. However, as the field of immunotherapy advances, researchers are increasingly recognizing the limitations of IgG in certain therapeutic contexts, such as targeting mucosal pathogens, eradicating solid tumors, or treating severe allergic conditions. This realization has sparked a surge of interest in non-IgG isotypes, specifically IgA, IgM, and IgE.
While these alternative isotypes offer unique effector functions and binding properties, bringing them to the clinic requires overcoming significant developmental hurdles. Chief among these are humanization and de-immunization. While the foundational concepts remain similar to IgG, transitioning these engineering processes to non-IgG projects introduces entirely new layers of structural and immunological complexity.
In this comprehensive guide, we will explore the critical differences between IgG and non-IgG therapeutic engineering, detailing the unique challenges and strategies involved in the humanization and de-immunization of IgA, IgM, and IgE antibodies.
The Core Concepts: Humanization and De-immunization
Before diving into isotype-specific nuances, it is crucial to establish baseline definitions for these two essential engineering processes.
-
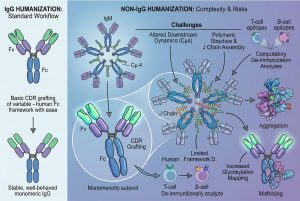
Humanization: This process involves modifying non-human antibodies (typically from mice or rabbits) so that they closely resemble human antibodies. The most common technique is Complementarity Determining Region (CDR) grafting, where the antigen-binding loops of the animal antibody are transplanted onto a human framework. The goal is to retain antigen specificity and affinity while reducing the antibody’s foreignness to the human immune system.
-
De-immunization: Even fully human or humanized antibodies can elicit an immune response, leading to the formation of Anti-Drug Antibodies (ADAs). De-immunization goes a step further by utilizing computational prediction and in vitro assays to identify and eliminate specific T-cell or B-cell epitopes within the antibody sequence. By mutating these immunogenic hotspots, researchers can significantly reduce the risk of clinical immunogenicity.
While the pharmaceutical industry has perfected these techniques for IgG, applying them to IgA, IgM, and IgE requires a fundamental shift in strategy due to their distinct biological and structural characteristics.
Why the Shift to Non-IgG Isotypes?
To understand the engineering challenges, we must first look at why researchers are turning to these complex molecules.
-
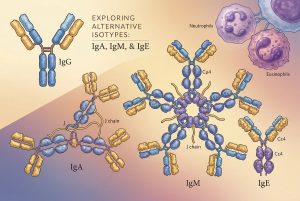
IgA (Immunoglobulin A): As the primary antibody of mucosal immunity, IgA is uniquely equipped to operate in the gastrointestinal and respiratory tracts. Furthermore, IgA interacts with the FcαRI (CD89) receptor on neutrophils and macrophages, triggering potent tumor-killing mechanisms that often surpass IgG-mediated cellular cytotoxicity.
-
IgM (Immunoglobulin M): IgM forms massive pentameric or hexameric structures, granting it up to ten antigen-binding sites. This provides unparalleled avidity, making IgM incredibly effective at binding repetitive target antigens on viral surfaces or cancer cells. It is also the most potent isotype for activating the classical complement cascade.
-
IgE (Immunoglobulin E): Though infamous for its role in allergies, IgE has a profoundly strong affinity for its receptor, FcεRI. In oncology, IgE therapeutics can recruit powerful immune cells, such as eosinophils and mast cells, to aggressively attack solid tumors that are otherwise resistant to IgG-based therapies.
Structural Complexities: How Non-IgG Differs from IgG
The primary reason humanization and de-immunization for non-IgG projects differ from IgG lies in the fundamental architecture of the molecules.
1. Additional Constant Domains
Unlike IgG, which contains three heavy chain constant domains (CH1, CH2, and CH3), both IgM and IgE possess a fourth domain (Cμ4 and Cε4, respectively). This extended structure alters the overall flexibility and spatial geometry of the antibody. When performing humanization, back-mutations in the framework regions (FRs) must account for these altered downstream structural dynamics to ensure the variable region does not collapse or lose its binding affinity.
2. Polymeric Assemblies and the J Chain
IgA and IgM are rarely simple monomers. IgA often circulates as a dimer, while IgM forms pentamers or hexamers. These multimeric forms are held together by disulfide bonds and a distinct polypeptide called the Joining (J) chain. Humanizing a murine IgM requires not only optimizing the variable regions but also ensuring that the engineered heavy chains can still correctly oligomerize with the J chain. A poorly designed framework graft can lead to steric hindrance, preventing proper pentamer formation and resulting in aggregated, non-functional proteins.
3. Extensive Glycosylation Profiles
Non-IgG isotypes are significantly more glycosylated than IgG. For example, IgM has five highly conserved N-linked glycosylation sites on its heavy chain, and IgA possesses complex O-linked glycans in its hinge region. During de-immunization, these glycans present a dual challenge:
-
The Shielding Effect: Glycans can shield underlying amino acid sequences from immune recognition. If an engineering step inadvertently alters a glycosylation site, a previously hidden immunogenic epitope might be exposed.
-
Neo-epitope Generation: Conversely, removing or altering glycans during humanization can create novel immunogenic hotspots. Predictive in silico models must be highly sophisticated to account for 3D glycan shielding when mapping T-cell epitopes for non-IgG isotypes.
Unique Challenges in Non-IgG Humanization
When transitioning a non-IgG project from discovery to lead optimization, scientists face several specialized hurdles.
Limited Human Framework Databases
For IgG humanization, researchers have access to massive databases of human variable gene repertoires, making it relatively straightforward to select an ideal human framework for CDR grafting. For non-IgG antibodies, the available repertoire data is less exhaustive. Matching the non-human sequence to a suitable human IgA, IgM, or IgE framework requires more rigorous computational modeling and often involves generating hybrid or chimeric intermediates to test functional stability before full humanization.
Managing Developability and Aggregation
One of the most critical factors distinguishing IgG from non-IgG projects is developability. Because IgM and IgA are multimeric and highly complex, they are incredibly prone to aggregation, misfolding, and poor expression yields during mammalian cell culture production. When humanizing these molecules, every amino acid substitution must be heavily scrutinized not just for its impact on binding, but for its impact on thermodynamic stability. Minor framework changes that an IgG molecule could easily tolerate might cause a humanized IgM pentamer to completely precipitate out of solution.
Advanced De-immunization Strategies for IgA, IgM, and IgE
Removing immunogenic risk from non-IgG therapeutics requires moving beyond basic sequence-based prediction.
3D Conformational Epitope Mapping: Because of the complex quaternary structures of multimeric IgM and dimeric IgA, linear sequence analysis is insufficient. An amino acid sequence that appears safe in a linear model might form a potent conformational epitope when folded into a hexameric structure. De-immunization of non-IgG antibodies heavily relies on advanced 3D structural modeling to identify spatial epitopes created at the interfaces of the varied domains.
Chimeric Engineering Approaches: In some complex projects, a stepwise approach is necessary. Researchers may first create an engineered chimeric antibody—for example, splicing the variable regions of an IgM onto an IgG constant backbone, or vice versa. This allows scientists to isolate the binding variables from the effector variables, enabling them to map and remove immunogenic hotspots systematically before reassembling the final therapeutic molecule.
Conclusion
The transition from IgG-centric drug development to the exploration of IgA, IgM, and IgE is unlocking exciting new frontiers in oncology, infectious diseases, and allergy treatments. However, unlocking this potential requires acknowledging that these molecules are not simply “IgG with a different tail.” Their multimeric structures, heavy glycosylation, and distinct molecular dynamics require highly specialized approaches to humanization and de-immunization. By utilizing advanced 3D modeling, careful framework selection, and rigorous developability assessments, developers can successfully transform non-human binders into safe, highly effective non-IgG therapeutics.
Accelerate Your Non-IgG Therapeutic Development
At Creative Biolabs, we understand the unique structural and immunological demands of alternative antibody isotypes. We offer a comprehensive suite of customized engineering solutions to help you overcome the hurdles of humanization, de-immunization, and developability, ensuring your lead candidates are optimized for clinical success.
Explore our specialized non-IgG engineering services:
Partner with us to navigate the complexities of non-IgG therapeutic development and bring your innovative treatments to the patients who need them.